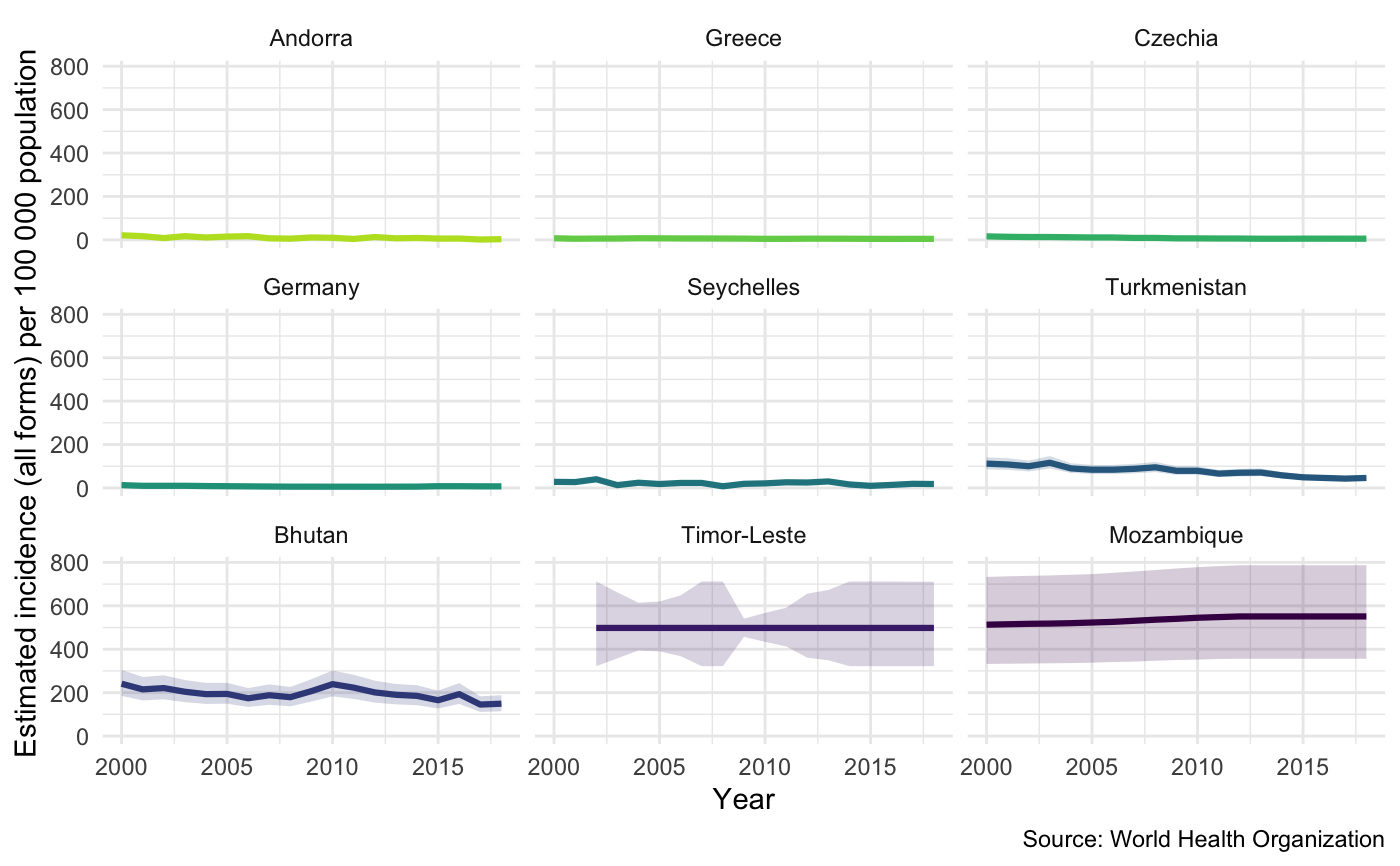
Plot measures of TB burden by country by specifying a metric from the TB burden data. Specify a country or vector of countries in order to plot them (by default plots all countries). Various other options are available for tuning the plot further.
plot_tb_burden( df = NULL, dict = NULL, metric = "e_inc_100k", metric_label = NULL, smooth = FALSE, conf = c("_lo", "_hi"), countries = NULL, years = NULL, compare_to_region = FALSE, facet = NULL, legend = "none", annual_change = FALSE, trans = "identity", scales = "fixed", interactive = FALSE, download_data = TRUE, save = TRUE, viridis_palette = "viridis", viridis_direction = -1, viridis_end = 0.9, verbose = FALSE, ... )
Arguments
| df | Dataframe of TB burden data, as sourced by |
|---|---|
| dict | A tibble of the data dictionary. See |
| metric | Character string specifying the metric to plot |
| metric_label | Character string specifying the metric label to use. |
| smooth | Logical, defaults to |
| conf | Character vector specifying the name variations to use to specify the upper
and lower confidence intervals. Defaults to c("_lo", "_hi"), if set to |
| countries | A character string specifying the countries to target. |
| years | Numeric vector of years. Defaults to |
| compare_to_region | Logical, defaults to |
| facet | Character string, the name of the variable to facet by. |
| legend | Character string, defaults to |
| annual_change | Logical, defaults to |
| trans | A character string specifying the transform to use on the specified metric. Defaults to no
transform ("identity"). Other options include log scaling ("log") and log base 10 scaling
("log10"). For a complete list of options see |
| scales | Character string, see ?ggplot2::facet_wrap for details. Defaults to "fixed", alternatives are "free_y", "free_x", or "free". |
| interactive | Logical, defaults to |
| download_data | Logical, defaults to |
| save | Logical, should the data be saved for reuse during the current R session. Defaults to
|
| viridis_palette | Character string indicating the |
| viridis_direction | Numeric, indicating the direction for the colour palette (1 or -1), defaults to -1.
See |
| viridis_end | Numeric between 0 and 1, defaults to 0.9. The end point of the viridis scale to use.
#' See |
| verbose | Logical, defaults to |
| ... | Additional arguments to pass to |
Value
A plot of TB Incidence Rates by Country
See also
get_tb_burden search_data_dict
Examples
## Get the WHO TB burden data and the data dictionary tb_burden <- get_tb_burden() dict <- get_data_dict() ## Get a random sample of 9 countries sample_countries <- sample(unique(tb_burden$country), 9) ## Plot incidence rates in these countries plot_tb_burden(df = tb_burden, dict = dict, facet = "country", countries = sample_countries)if (FALSE) { ## Plot smoothed incidence rates in these countries plot_tb_burden( df = tb_burden, dict = dict, facet = "country", smooth = TRUE, countries = sample_countries ) ## Use data caching to plot incidence rates with free y scales plot_tb_burden(facet = "country", countries = sample_countries, scales = "free_y") ## Plot annual percentage change in incidence rates in selected countries plot_tb_burden( df = tb_burden, dict = dict, facet = "country", scales = "free_y", countries = sample_countries, annual_change = TRUE, conf = NULL ) ## Find variables relating to mortality in the WHO dataset search_data_dict(def = "mortality") ## Plot mortality rates (exc HIV) - without progress messages plot_tb_burden( metric = "e_mort_exc_tbhiv_100k", facet = "country", countries = sample_countries, scales = "free_y" ) }